Multiscale Kinetic Architectures of the IFN-γ Interactome
The goal of the project was to describe and map quantitatively time-dependent profile of Interferon-gamma (IFN-γ) and its role in macrophage activation. To achieve it, I have employed object-oriented, equation-based Modelica language. The work moved beyond standard ODE solvers and succeeded to create a modular, compartmentalized framework capable of simulating non-linear feedback loops and inter-cellular interactions in the immune system.
Engineering of Kinetic Models
The effort focused on the IFN-γ signaling cascade, constructing models to simulate the transition of macrophages from a patrolling state to an activated state. This transition is triggered by a dual-pathway mechanism: non-specific bacterial signals (LPS) and cytokine-driven pathways involving TNF and IFN-γ.
Design of the Custom ImSys Library
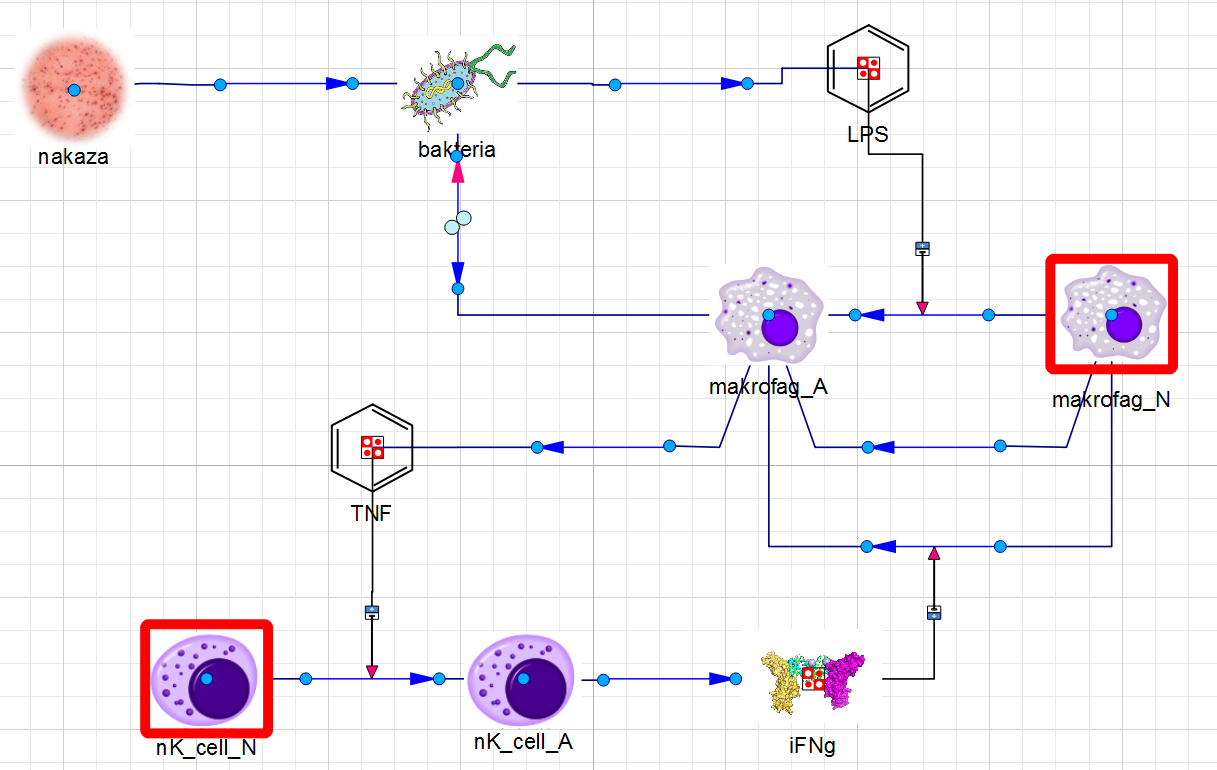
Implemented kinetics scheme
I developed ImSys, a specialized bio-computational library that facilitates the construction of complex kinetic models. Unlike generic tools, ImSys handles biological specificities such as autocatalytic cell proliferation and resource-constrained growth. Through a graphical interface, these components allow for the elegant investigation of process dependencies or the rapid reconfiguration of system components.
Simulations and Physiological Insights
Using OpenModelica and Wolfram SystemModeler, I conducted sensitivity analyses on key kinetic parameters. The simulations revealed critical behaviors in the immune response:
- Signal Amplification: Quantifying how IFN-γ production by NK cells significantly strengthens the macrophage response.
- Stability & Oscillations: Identifying parameter sets that lead to excessive activation or periodic inflammatory surges—phenomena characteristic of autoimmune conditions like psoriasis.
The architecture was designed for verification against real-world clinical data, providing a robust platform for testing therapeutic hypotheses.
Technical Evolution (Legacy & Future)
While originally realized using the Modelica platform, the underlying ImSys logic remains platform-agnostic. The mathematical models are fully compatible with modern FMI (Functional Mock-up Interface) standards, allowing for seamless integration into contemporary Python or Julia-based research environments.