Projects / Bioinformatics / Monoclonal Antibodies
Comprehensive Biological ETL-EDA Workflow
Systematic Domain Architecture Analysis of mAbs
This bioinformatics pipeline categorizes Monoclonal Antibodies (mAbs) based on their domain composition and critical structural characteristics, such as Complementarity-Determining Regions (CDRs) and disulfide bridges. It provides a rigorous foundation for targeted modifications, novelty assessment, and rational design
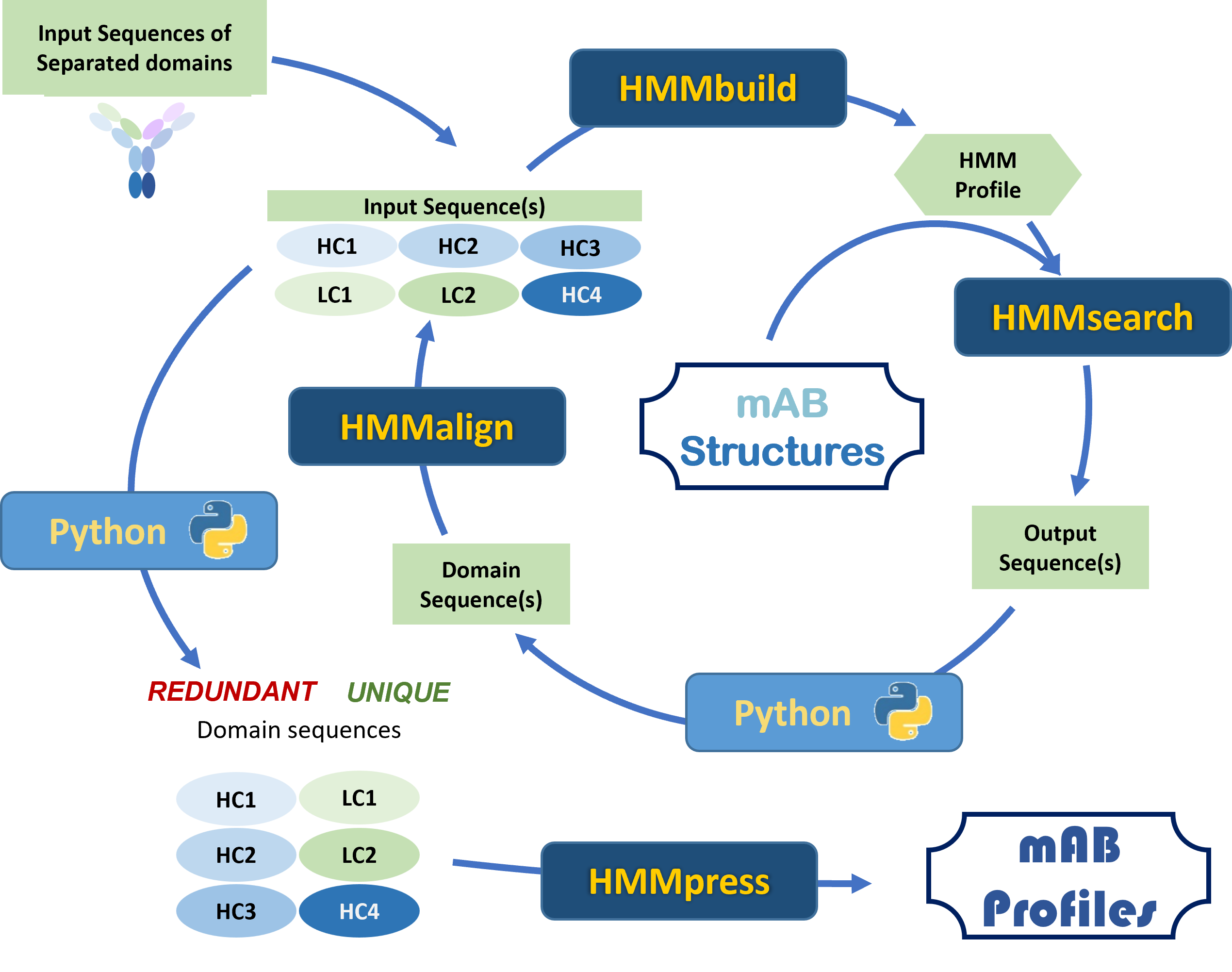
The workflow orchestrated via Python integrates HMMbuild, HMMsearch, HMMalign, and HMMpress.
The system enables a thorough exploration of unknown mAb query sequences
- Rapid Topology identification, detection of structural components and domain boundaries.
- Mapping of residues interacting with antigens and location of potential glycosylation sites.
- Prediction of CDR conformations.
I implemented this high-throughput pipeline to process the full library of antibody sequences derived from the Protein Data Bank (PDB). The system creates specialized profile databases for individual mAb domains, transforming raw structural data into a searchable, HMM-based repository of domain architectures.