Projects / Bioinformatics / mAB Design
Web App / Bioinformatics
Interactive, Template-Guided Platform for mABs (re)construction
This web-based interface utilizes an integrated backend workflow that combines HMMER-based domain profiling with MODELLER. It enables users to create adjustable models of monoclonal antibodies (mAbs) by mapping their specific domain architecture and identifying key regions that interact with antigens
Project Overview
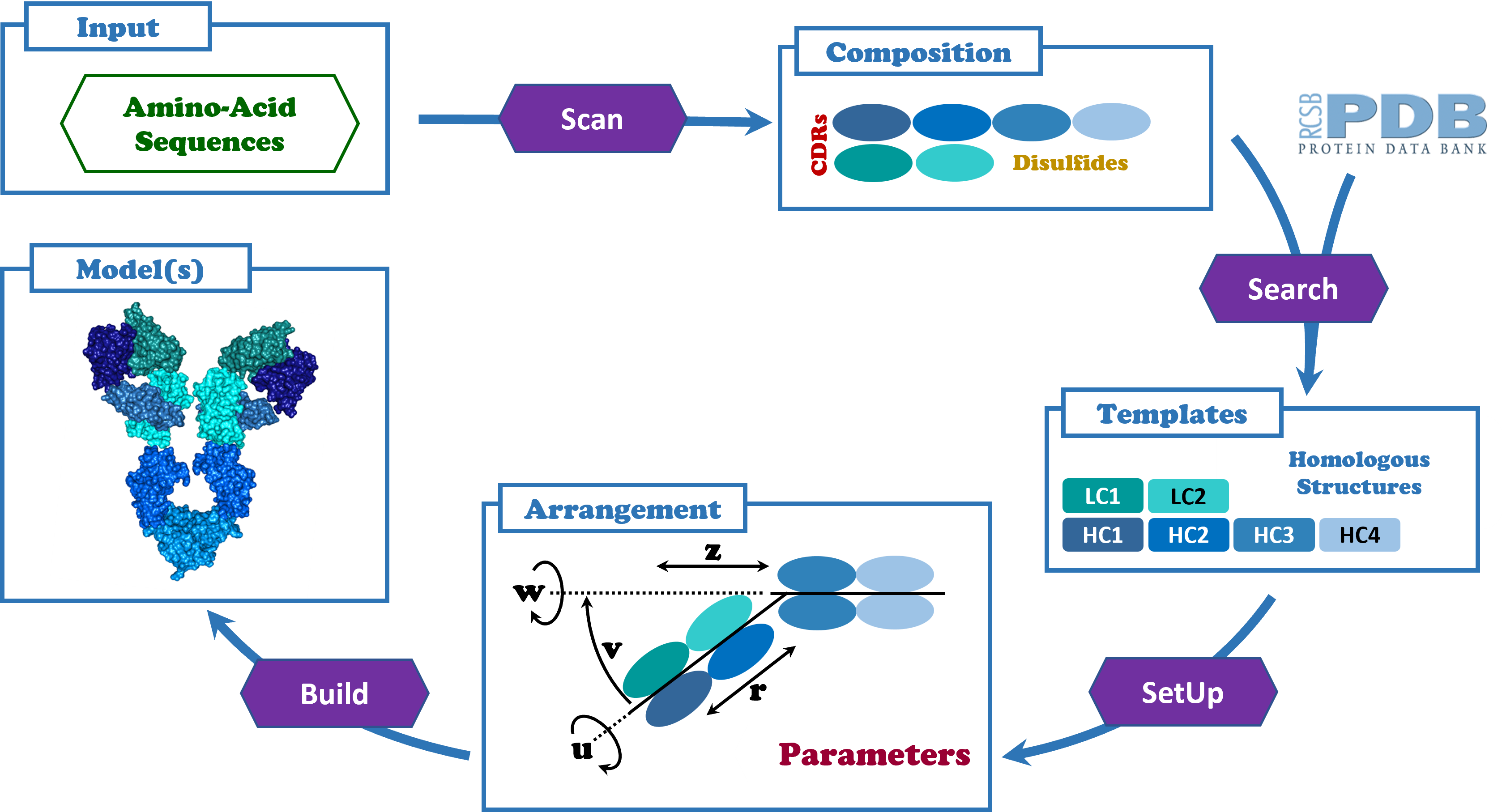
Workflow Pipeline
- Scan input amino-acid sequence(s) to detect domains, Complementary-Determining Regions (CDRs), and disulfide bonds.
- Support implemented antibody types with a guided setup flow.
- Search PDB for homologous structures per antibody domain.
- Select structural templates independently for each domain.
- Define 3D arrangement through translation and rotation parameters.
- Build final model structures for each selected template and parameter set.
Engineering Notes
The current release focuses on a robust baseline pipeline with constrained design choices. The architecture is prepared for iterative expansion of antibody classes, smarter template ranking, and richer model validation layers.